 |
|
|
[Sponsors] | |||||
|
|
|
#1 |
|
Member
Krishna
Join Date: Apr 2012
Posts: 54
Rep Power: 14  |
Hi,
I used example file therm.dat but while simulation i am getting below error on nagative cp for c6H3 species. Error message: application called MPI_Abort(MPI_COMM_WORLD, 1) - process 0 JOB ABORT invoked by rank 3: ERROR: species 39 (C6H3) was found to have a negative Cp, please revise thermo data for this species in therm.dat c6h3 from them.dat C6H3 C 6H 3 0 0 298.000 5000.000 1000.00 1 0.58188343E+01 0.27933408E-01-0.17825427E-04 0.53702536E-08-0.61707627E-12 2 0.85188250E+05-0.92147827E+00 0.11790619E+01 0.55547360E-01-0.73076168E-04 3 0.52076736E-07-0.15046964E-10 0.85647312E+05 0.19179199E+02 4 Please somebody suggest me, which value represents cp in therm.dat. Thank you in advance. Best Regards, Krishna |
|
|
|

|
|
|
|
|
#2 |
|
Senior Member
Sameera Wijeyakulasuriya
Join Date: Jan 2016
Location: Convergent Science, Madison WI
Posts: 117
Rep Power: 10  |
Hi,
You can correct for this in CONVERGE Studio. Simply load the therm.dat in to Studio and go to Gas Simulation --> Gas thermodynamic data From the list of gas phase species, select C6h3 and use the plot button to open the plotting tool. Select "Specific Heat Capacity Cp" for Y-variable. You should be able to figure out at what temperature (X-variable), Cp goes negative. Let's say it starting to go negative at 3600 K. Then open therm.dat in a text editor outside of Studio and change the max temperature for this species to be 3500 K. What you are essentially doing here is to limit the maximum temperature for this species such that Cp is positive at max temperature. If the cell temperature goes beyond 3500 K in the simulation, CONVERGE will keep using the Cp at 3500 K for higher temperatures. |
|
|
|

|
|
|
|
|
#3 |
|
Member
Krishna
Join Date: Apr 2012
Posts: 54
Rep Power: 14  |
Thank you so much for your response.
|
|
|
|

|
|
|
|
|
#4 |
|
New Member
Alirezak
Join Date: Feb 2017
Posts: 24
Rep Power: 9  |
Dear Sir,
I have same error in converge and I checked Cp graph but there isnt any negative area for Cp the error is " WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C4H4O. Low Cp/R = 20.1433, High Cp/R = 20.525" and C4H4O C 4H 4O 1 G 300.00 4000.00 1000.00 1 .135912211E+02 .987450910E-02-.345325238E-05 .544429808E-09-.319349896E-13 2 -.115639534E+05-.542513644E+02-.750030326E+01 .699899868E-01-.695485649E-04 3 .333968809E-07-.619467883E-11-.541634292E+04 .550218432E+02 4 |
|
|
|

|
|
|
|
|
#5 |
|
Senior Member
Sameera Wijeyakulasuriya
Join Date: Jan 2016
Location: Convergent Science, Madison WI
Posts: 117
Rep Power: 10  |
Hello,
The 14 coefficients you see in therm.dat for a species defines two polynomials. 7 coefficients define the property at high temperature and the other 7 at low temperature. The two polynomials have to be continuous at the mid temperature value specified in therm.dat. This warning states that it is not the case for this species and asking you to correct it. Please email your therm.dat and mech.dat to Support@convergecfd.com so that we can correct this discrepancy for you. Thanks, |
|
|
|

|
|
|
|
|
#6 |
|
New Member
Alirezak
Join Date: Feb 2017
Posts: 24
Rep Power: 9  |
Dear Sir,
I would like to inform you that I dont have official email if there is no problem I send the file to you all errors in the mechanism WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C4H4O. Low Cp/R = 20.1433, High Cp/R = 20.525 WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C5H4O2. Low Cp/R = 24.1273, High Cp/R = 24.5862 WARNING: Therm.dat: Mismatch for H/RT at common temperatures. Species = C5H4O2. Low H/RT = -4.68497, High H/RT = -4.74867 WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C5H8O. Low Cp/R = 29.1639, High Cp/R = 29.7796 WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C5H8O4. Low Cp/R = 36.4342, High Cp/R = 37.1157 WARNING: Therm.dat: Mismatch for H/RT at common temperatures. Species = DMF. Low H/RT = 1.66077, High H/RT = 1.6805 WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C6H10O5. Low Cp/R = 45.7391, High Cp/R = 46.6112 WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = DIFENET. Low Cp/R = 49.9648, High Cp/R = 50.9154 WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C4H3O. Low Cp/R = 18.1735, High Cp/R = 18.5101 WARNING: Therm.dat: Mismatch for Cp/R at common temperature. Species = C5EN-OOQOOH-35. Low Cp/R = 38.9238, High Cp/R = 39.6945 WARNING: Therm.dat: Mismatch for S/R at common temperatures. Species = ZMSTEAOOH. Low S/R = 189.435, High S/R = 260.203 |
|
|
|

|
|
|
|
|
#7 |
|
New Member
Alirezak
Join Date: Feb 2017
Posts: 24
Rep Power: 9  |
I will try it in matlab
|
|
|
|

|
|
|
|
|
#8 |
|
New Member
Alirezak
Join Date: Feb 2017
Posts: 24
Rep Power: 9  |
Dear Sir/Madam,
I would like to inform you that during reduction a mechanism five isomers created in my reaction mechanism where it produced an error in transport data during 1D simulation, could you please explain how can i creating transport data for lumped isomers? the error is: transport data is missing for species 116, ISM001 transport data is missing for species 117, ISM002 transport data is missing for species 118, ISM003 transport data is missing for species 119, ISM004 transport data is missing for species 120, ISM005 Bests |
|
|
|

|
|
|
|
|
#9 |
|
New Member
Aditya Raman
Join Date: Jul 2020
Posts: 10
Rep Power: 5  |
Hi Alireza,
The transport data for lumped isomers should be written along with the mechanism data. Can you describe what you were trying to do and when the following error happens? Regards, ------------------------------------ Aditya Raman, PhD Research Engineer, Applications Convergent Science ------------------------------------ |
|
|
|

|
|
|
|
|
#10 |
|
New Member
Alirezak
Join Date: Feb 2017
Posts: 24
Rep Power: 9  |
thank you for your answer,
I would like to inform you that I am trying to reduce Zhang2020 biodiesel mechanism i converge, with DRGEPSA and isomer lumping, the final mechanism consists of lumped isomers with appropriate data in therm_lump.dat file, but there is no information related to Isomers in transport.dat because the software doesn't generate transport.dat for reduction mechanism. |
|
|
|

|
|
|
|
|
#11 |
|
New Member
Aditya Raman
Join Date: Jul 2020
Posts: 10
Rep Power: 5  |
You would have to check on the Enable Transport option under Mechanism Reduction > Target Species and tolerances > Isomer Lumping tab in Studio.
After this you will have to include your transport.dat input file and the mechanism reduction tool should write out your transport_lumped.dat file. Regards, ------------------------------------ Aditya Raman, PhD Research Engineer, Applications Convergent Science ------------------------------------ |
|
|
|

|
|
|
|
|
#12 |
|
New Member
Alirezak
Join Date: Feb 2017
Posts: 24
Rep Power: 9  |
thank you for your reply, according to attached figure there is no option for transport lumping in CONVERGE v2.4
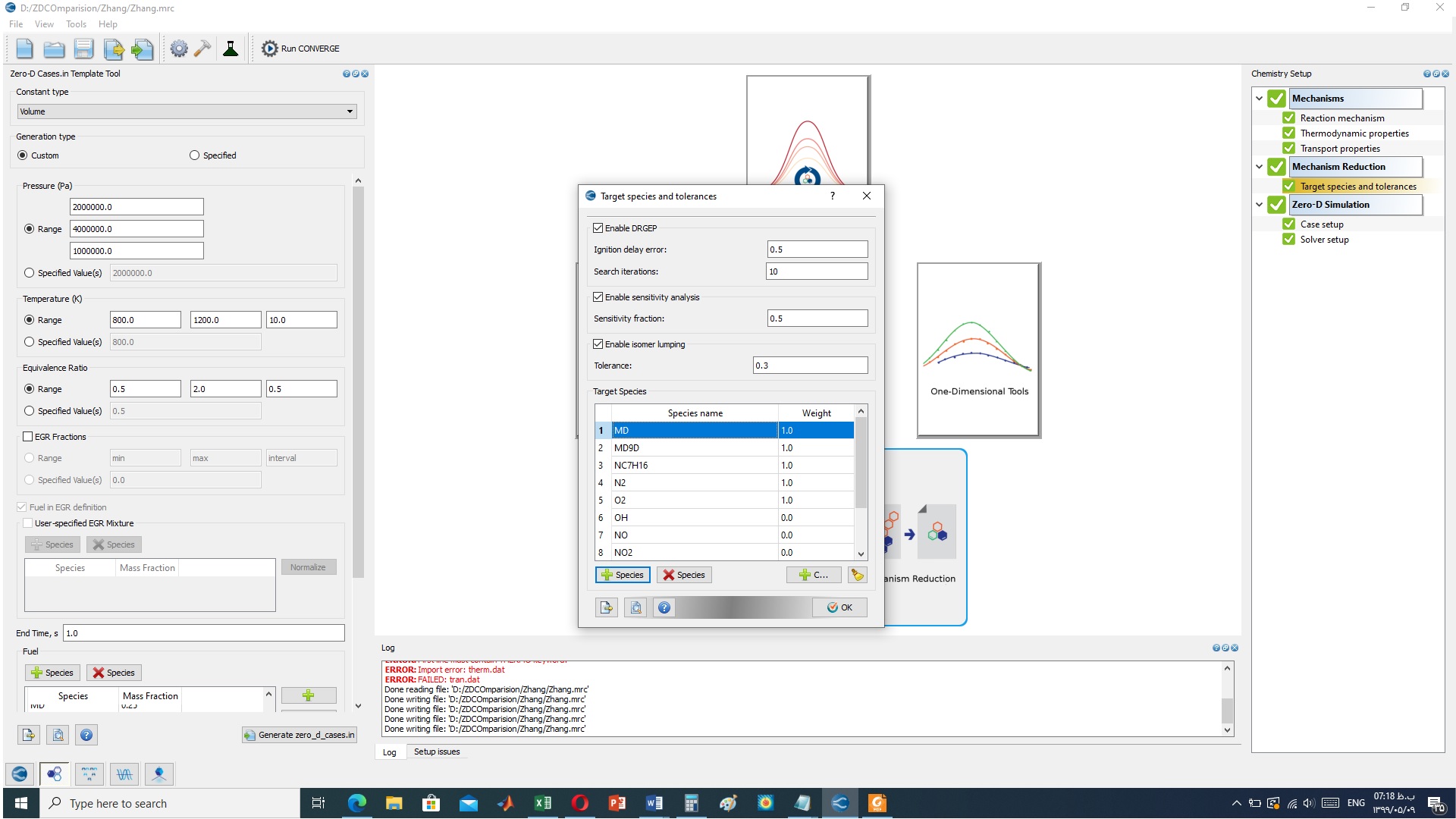 bests |
|
|
|

|
|
|
|
|
#13 |
|
New Member
Aditya Raman
Join Date: Jul 2020
Posts: 10
Rep Power: 5  |
Hello Alireza,
Which version of Studio are you currently using? Try updating it to the most recent one. You will see an option to enable transport, similar to the attached figure. If you have transport.dat file in the directory the isomer lumping should also read that and write out lumped transport data. Regards, ------------------------------------ Aditya Raman, PhD Research Engineer, Applications Convergent Science ------------------------------------ |
|
|
|

|
|
|
|
|
#14 |
|
New Member
Aditya Raman
Join Date: Jul 2020
Posts: 10
Rep Power: 5  |
If you are still having the same issue, please feel free to reach out to us at support@convergecfd.com (US); supportEU@convergecfd.com (EU) or support.in@convergecfd.com (India) with your case setup. We would be happy to have a look at this.
Regards, ------------------------------------ Aditya Raman, PhD Research Engineer, Applications Convergent Science ------------------------------------ |
|
|
|

|
|
 |
|
|
 Similar Threads
Similar Threads
|
||||
| Thread | Thread Starter | Forum | Replies | Last Post |
| [Other] mesh airfoil NACA0012 | anand_30 | OpenFOAM Meshing & Mesh Conversion | 13 | March 7, 2022 17:22 |
| [blockMesh] non-orthogonal faces and incorrect orientation? | nennbs | OpenFOAM Meshing & Mesh Conversion | 7 | April 17, 2013 05:42 |
| [blockMesh] error message with modeling a cube with a hold at the center | hsingtzu | OpenFOAM Meshing & Mesh Conversion | 2 | March 14, 2012 09:56 |
| [blockMesh] BlockMesh FOAM warning | gaottino | OpenFOAM Meshing & Mesh Conversion | 7 | July 19, 2010 14:11 |
| [blockMesh] Axisymmetrical mesh | Rasmus Gjesing (Gjesing) | OpenFOAM Meshing & Mesh Conversion | 10 | April 2, 2007 14:00 |